SUIT-001 O2 ce-pce D004: Difference between revisions
From Bioblast
No edit summary |
No edit summary |
||
| Line 51: | Line 51: | ||
== Compare SUIT protocols == | == Compare SUIT protocols == | ||
::::* [[ | ::::* [[SUIT-2_O2_pce_D07a]] (RP2): Harmonized SUIT protocol for PBMC and PLT cells. | ||
::::* [[SUIT-004]]: Short version of RP1; without G, F and Gp. | ::::* [[SUIT-004]]: Short version of RP1; without G, F and Gp. | ||
Revision as of 13:45, 21 January 2019
Description
Abbreviation: NSGp(PGM)
Reference: A Reference protocol RP1 specific for PBMC and PTL cells - SUIT-001
SUIT number: D004_ce1;1Dig;1PM;2D;2c;3U;4G;5S;7Rot;8Gp;9Ama;10AsTm;11Azd
O2k-Application: O2
- MitoPedia: SUIT - SUIT reference protocol RP1 for PBMCs and PLTs
- SUIT-category: NSGp(PGM)
- SUIT protocol pattern: orthogonal 1PM;2D;2c;3U;4G;5S;7Rot;8Gp
- Reference protocol 1 specifically for PBMCs and PLTs.
- The main difference between the RP1 for PBMCs and PLTs and RP1 for other mtprep SUIT RP1 is the absence of Oct in the case of blood cells.
References
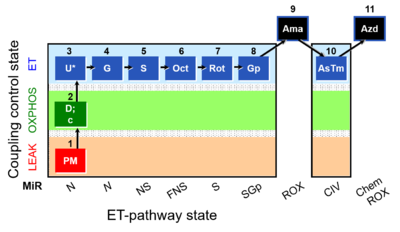
Steps and respiratory states
| Step | State | Pathway | Q-junction | Comment - Events (E) and Marks (M) |
|---|---|---|---|---|
| 1PM | PML(n) | N | CI | 1PM
|
| 2D | PMP | N | CI | 1PM;2D
|
| 2c | PMcP | N | CI | 1PM;2D;2c
|
| 3U | PME | N | CI | 1PM;2D;3U
|
| 4G | PGME | N | CI | 1PM;2D;3U;4G
|
| 5S | PGMSE | NS | CI&II | 1PM;2D;3U;4G;5S
|
| 6Oct | OctPGMSE | FNS | FAO&CI&II | 1PM;2D;3U;4G;5S;6Oct
|
| 7Rot | SE | S | CII | 1PM;2D;3U;4G;5S;6Oct;7Rot
|
| 8Gp | SGpE | SGp | CII&GpDH | 1PM;2D;3U;4G;5S;6Oct;7Rot;8Gp
|
| 9Ama | ROX | 1PM;2D;3U;4G;5S;6Oct;7Rot;8Gp;9Ama
|
| Step | Respiratory state | Pathway control | ET-Complex | Comment |
|---|---|---|---|---|
| ## AsTm | AsTmE | CIV | CIV | |
| ## Azd | CHB |
- Bioblast links: SUIT protocols - >>>>>>> - Click on [Expand] or [Collapse] - >>>>>>>
- Coupling control
- Pathway control
- Main fuel substrates
- » Glutamate, G
- » Glycerophosphate, Gp
- » Malate, M
- » Octanoylcarnitine, Oct
- » Pyruvate, P
- » Succinate, S
- Main fuel substrates
- Glossary
Strengths and limitations
- SUIT-001 in combination with SUIT-002 provides a common reference for comparison of respiratory control in a large variety of species, tissues and cell types. Both SUIT protocols provide a mitochondrial mapping which allows:
- 1. to obtain reference values.
- 2. to evaluate mitochondrial physiological diversity, generating a mt-database on comparative mitochondrial physiology.
- 3. to screen specific defects.
- SUIT-001 and SUIT-002 are used in the MitoFit Proficiency test for inter-individual and inter-laboratory reproducibility quality control.
- A succinate concentration of >10 mM may be required for saturating SE capacity.
- + Linear coupling control (L-P-E) with N substrates (PM).
- + Pathway control in ET state (N, NS, FNS, S and SGp pathways).
- + Harmonization with SUIT-002 allows to perform both SUIT protocols in parallel. The cross-linked respiratory states can be statically used as repeat measurements.
- + In blood cells, SUIT-001 and SUIT-002 harmonize in: ROUTINE (ce1), SGpE (8Gp in SUIT-001 and 10Rot in SUIT-002) and Complex IV.
- + Harmonization with many SUIT protocols (up to step 5S).
- + This protocol can be extended with the Complex IV module.
- - The substrate combination for maximum ET capacity (FNSGP) is not obtained in SUIT-001 in favour of measuring SE.
- - Oct is not added in ET state.
Compare SUIT protocols
- SUIT-2_O2_pce_D07a (RP2): Harmonized SUIT protocol for PBMC and PLT cells.
MitoPedia concepts:
SUIT protocol,
SUIT A,
Find
MitoPedia methods:
Respirometry
Labels:
SUIT-001