Difference between revisions of "MitoEAGLE Working Groups"
From Bioblast
Beno Marija (talk | contribs) |
Beno Marija (talk | contribs) |
||
| Line 1: | Line 1: | ||
{{MITOEAGLE}} | {{MITOEAGLE}} | ||
[[File:MITOEAGLE Working groups.jpg|right|400px| | [[File:MITOEAGLE Working groups.jpg|right|400px|MitoEAGLE Working Groups]] | ||
__TOC__ | __TOC__ | ||
[[File:MITOEAGLE concept.jpg|right|400px| | [[File:MITOEAGLE concept.jpg|right|400px|MitoEAGLE concept]] | ||
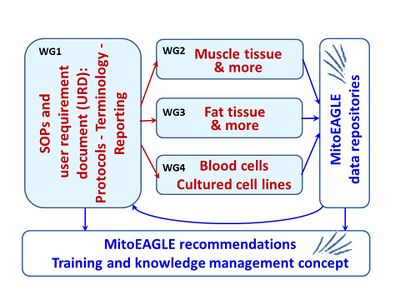
:::: Four Working Groups (WG) interact closely: WG1 elaborates the overall strategy, with input into Working Groups 2-4 specializing on various types of tissue and cells. Feeding the data repositories, WG2-4 will implement the strategies, providing feedback to WG1, with feedforward in optimization cycles. These will lead to summaries of recommendations and a ''Training and knowledge management concept''. | :::: Four Working Groups (WG) interact closely: WG1 elaborates the overall strategy, with input into Working Groups 2-4 specializing on various types of tissue and cells. Feeding the data repositories, WG2-4 will implement the strategies, providing feedback to WG1, with feedforward in optimization cycles. These will lead to summaries of recommendations and a ''Training and knowledge management concept''. | ||
| Line 12: | Line 12: | ||
::: '''WG2''' | ::: '''WG2''' | ||
:::: ''' | :::: '''MitoEAGLE data repository on muscle tissues''' | ||
::::» [[WG2 MITOEAGLE data: muscle |WG2 Action]] | ::::» [[WG2 MITOEAGLE data: muscle |WG2 Action]] | ||
::::» [[MITOEAGLE data: muscle |WG2 Project application]] | ::::» [[MITOEAGLE data: muscle |WG2 Project application]] | ||
::: '''WG3''' | ::: '''WG3''' | ||
:::: ''' | :::: '''MitoEAGLE data repository on fat, neuronal and liver tissues''' | ||
::::» [[WG3 MITOEAGLE data: adipose, liver, neuronal |WG3 Action]] | ::::» [[WG3 MITOEAGLE data: adipose, liver, neuronal |WG3 Action]] | ||
::::» [[MITOEAGLE data: fat |WG3 Project application]] | ::::» [[MITOEAGLE data: fat |WG3 Project application]] | ||
::: '''WG4''' | ::: '''WG4''' | ||
:::: ''' | :::: '''MitoEAGLE data repository for blood cells and cultured cells''' | ||
::::» [[WG4 MITOEAGLE data: blood and cultured cells |WG4 Action]] | ::::» [[WG4 MITOEAGLE data: blood and cultured cells |WG4 Action]] | ||
::::» [[MITOEAGLE data: blood and cultured cells |WG4 Project application]] | ::::» [[MITOEAGLE data: blood and cultured cells |WG4 Project application]] | ||
::: ''' | ::: '''MitoEAGLE data repositories''' (from WG2-4) | ||
::::# Deposit post-hoc datasets related to articles already published on topics of WG 2-4 by Consortium members on locations accessible for the Consortium. | ::::# Deposit post-hoc datasets related to articles already published on topics of WG 2-4 by Consortium members on locations accessible for the Consortium. | ||
::::# Connect published articles to deposited datasets, using generally accessible tools such as PubMed Commons (http://www.ncbi.nlm.nih.gov/pubmedcommons/). | ::::# Connect published articles to deposited datasets, using generally accessible tools such as PubMed Commons (http://www.ncbi.nlm.nih.gov/pubmedcommons/). | ||
Revision as of 16:06, 4 October 2017
| News and Events | Working Groups | Short-Term Scientific Missions | Management Committee | Members |
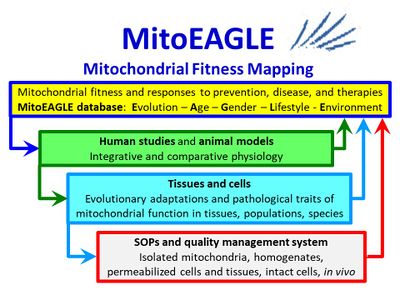
COST Action CA15203 (2016-2021): MitoEAGLE
Evolution-Age-Gender-Lifestyle-Environment: mitochondrial fitness mapping
MitoEAGLE Working Groups
- Four Working Groups (WG) interact closely: WG1 elaborates the overall strategy, with input into Working Groups 2-4 specializing on various types of tissue and cells. Feeding the data repositories, WG2-4 will implement the strategies, providing feedback to WG1, with feedforward in optimization cycles. These will lead to summaries of recommendations and a Training and knowledge management concept.
- WG1
- SOPs and user requirement document (URD): Protocols - Terminology - Reporting
- » WG1 Action
- » WG1 Project application
- WG1
- WG2
- MitoEAGLE data repository on muscle tissues
- » WG2 Action
- » WG2 Project application
- WG2
- WG3
- MitoEAGLE data repository on fat, neuronal and liver tissues
- » WG3 Action
- » WG3 Project application
- WG3
- WG4
- MitoEAGLE data repository for blood cells and cultured cells
- » WG4 Action
- » WG4 Project application
- WG4
- MitoEAGLE data repositories (from WG2-4)
- Deposit post-hoc datasets related to articles already published on topics of WG 2-4 by Consortium members on locations accessible for the Consortium.
- Connect published articles to deposited datasets, using generally accessible tools such as PubMed Commons (http://www.ncbi.nlm.nih.gov/pubmedcommons/).
- Along with development of SOPs, implement advance public deposition of protocols (Begley, Ioannidis 2015).
- Deposit datasets with submitted manuscripts and finally connect these using tools such as PubMed Commons.
- MitoEAGLE data repositories (from WG2-4)