Pesta 2012 Methods Mol Biol: Difference between revisions
No edit summary |
No edit summary |
||
| Line 24: | Line 24: | ||
|pathways=F, N, S, NS, ROX | |pathways=F, N, S, NS, ROX | ||
|instruments=Oxygraph-2k, TIP2k, O2k-Protocol | |instruments=Oxygraph-2k, TIP2k, O2k-Protocol | ||
|additional=MitoPathways, O2k-chemicals and media, Mt-preparations, O2k-Demo, O2k-Core, 1OctM;2D;3G;4S;5U;6Rot- | |additional=MitoPathways, O2k-chemicals and media, Mt-preparations, O2k-Demo, O2k-Core, 1OctM;2D;3G;4S;5U;6Rot-, SUIT-017 | ||
}} | }} | ||
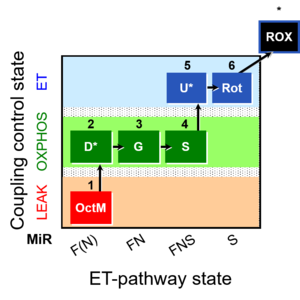
[[File:1OctM;2D;3G;4S;5U;6Rot-.jpg|right|300px|link=1OctM;2D;3G;4S;5U;6Rot- |1OctM;2D;3G;4S;5U;6Rot-]] | [[File:1OctM;2D;3G;4S;5U;6Rot-.jpg|right|300px|link=1OctM;2D;3G;4S;5U;6Rot- |1OctM;2D;3G;4S;5U;6Rot-]] | ||
== SUIT protocol == | == SUIT protocol == | ||
****: [[1OctM;2D;3G;4S;5U;6Rot-]] | ****: [[SUIT-017 |1OctM;2D;3G;4S;5U;6Rot-]] | ||
'''»''' [[MitoPedia: Respiratory states]] [[Image:P.jpg|link=OXPHOS-capacity|OXPHOS]] [[Image:R.jpg|link=ROUTINE respiration|ROUTINE]] [[Image:E.jpg|link=ET-capacity|ET-capacity]] [[Image:L.jpg|link=LEAK respiration|LEAK]] [[Image:ROX.jpg|link=Residual oxygen consumption|ROX]] | '''»''' [[MitoPedia: Respiratory states]] [[Image:P.jpg|link=OXPHOS-capacity|OXPHOS]] [[Image:R.jpg|link=ROUTINE respiration|ROUTINE]] [[Image:E.jpg|link=ET-capacity|ET-capacity]] [[Image:L.jpg|link=LEAK respiration|LEAK]] [[Image:ROX.jpg|link=Residual oxygen consumption|ROX]] | ||
Revision as of 11:55, 15 January 2019
| Pesta D, Gnaiger E (2012) High-Resolution Respirometry. OXPHOS protocols for human cells and permeabilized fibres from small biopsies of human muscle. Methods Mol Biol 810:25-58. |
Pesta D, Gnaiger E (2012) Methods Mol Biol
Abstract:
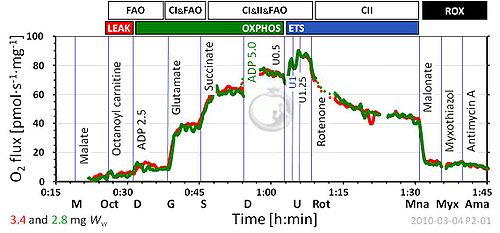
Protocols for High-Resolution Respirometry of intact cells, permeabilized cells, and permeabilized muscle fibres offer sensitive diagnostic tests of integrated mitochondrial function using standard cell culture techniques and small needle biopsies of muscle. Substrate-uncoupler-inhibitor titration (SUIT) protocols for analysis of oxidative phosphorylation improve our understanding of mitochondrial respiratory control and the pathophysiology of mitochondrial diseases. Respiratory states are defined in functional terms to account for the network of metabolic interactions in complex SUIT protols with stepwise modulation of coupling and substrate control. A regulated degree of intrinsic uncoupling is a hallmark of oxidative phosphorylation, whereas pathological and toxicological dyscoupling is evaluated as a mitochondrial defect. The noncoupled state of maximum respiration is experimentally induced by titration of established uncouplers (FCCP, DNP), to decrease the proton gradient across the mitochondrial inner membrane and measure the capacity of the electron transfer-pathway (ET-pathway, open-circuit operation of respiration). Intrinsic uncoupling and dyscoupling are evaluated as the flux control ratio between non-phosphorylating LEAK respiration (electron flow coupled to proton pumping to compensate for proton leaks) and ET-capacity. If OXPHOS-capacity (maximally ADP stimulated oxygen flux) is less than ET-capacity, the phosphorylation system contributes to flux control. Physiological NS-substrate combinations support maximum ET-pathway and OXPHOS capacities, due to the additive effect of multiple electron supply pathways converging at the Q-junction. Substrate control with electron entry separately through Complex I (pyruvate&malate or glutamate&malate) or Complex II (succinate&rotenone) restricts ET-capacity and artificially enhances flux control upstream of the Q-cycle, providing diagnostic information on specific branches of the ET-pathway. Oxygen levels are maintained above air saturation in protocols with permeabilized muscle fibres to avoid experimental oxygen limitation of respiration.
Standardized two-point calibration of the polarographic oxygen sensor (static sensor calibration), calibration of the sensor response time (dynamic sensor calibration), and evaluation of instrumental background oxygen flux (systemic flux compensation) provide the unique experimental basis for high accuracy of quantitative results and quality control in High-Resolution Respirometry.
Terminology and abbreviations edited according to MitoEAGLE recommendations (Gnaiger E).*]] • Keywords: Substrate-uncoupler-inhibitor titration, Human vastus lateralis, Needle biopsy, HEK, HPMC, HUVEC, Fibroblasts, Routine respiration, Oxidative phosphorylation, Q-junction, Pyruvate, Glutamate, Malate, Succinate, Leak, Coupling control, Uncoupling, Oxygraph, Oxygen flux, Residual oxygen consumption, Instrumental background
• O2k-Network Lab: AT Innsbruck Gnaiger E, AT Innsbruck Oroboros, AT Innsbruck MitoCom
Labels: MiParea: Respiration, Instruments;methods, mt-Biogenesis;mt-density, Comparative MiP;environmental MiP, Exercise physiology;nutrition;life style, mt-Medicine
Organism: Human
Tissue;cell: Skeletal muscle, Other cell lines, HEK, Fibroblast, HUVEC
Preparation: Intact cells, Permeabilized cells, Permeabilized tissue
Enzyme: Marker enzyme
Regulation: ADP, Coupling efficiency;uncoupling, Cyt c, Flux control, Oxygen kinetics, Substrate, Threshold;excess capacity, Fatty acid
Coupling state: LEAK, ROUTINE, OXPHOS, ET
Pathway: F, N, S, NS, ROX
HRR: Oxygraph-2k, TIP2k, O2k-Protocol
MitoPathways, O2k-chemicals and media, Mt-preparations, O2k-Demo, O2k-Core, 1OctM;2D;3G;4S;5U;6Rot-, SUIT-017
SUIT protocol
» MitoPedia: Respiratory states ![]()
![]()
![]()
![]()

Correction
- Tab. 2: ET-capacity of HEK 293 cells is 47 ± 7 (the value of 14 ± 2 given in the table is that of ROUTINE respiration).
- Fig. 7a: 'Myothiazol' should be corrected to 'Myxothiazol'.
Update
- Tab. 1: Malate was generally used at 2 mM final concentration. New evaluations of SUIT protocols analyze the inhibitory effect of malate on CII-linked respiration, and the possibility to compensate for this by increasing succinate concentrations >10 mM.
- » More details: Malate
- New terminology (Gnaiger 2014 MitoPathways): (i) 'ETF+CI' was used to indicate fatty acid oxidation (FAO), when electron-transferring flavoprotein complex (CETF) and Complex I (CI) are stimulated simultaneously (octanoylcarnitine&malate). Since this combined action is a general reqirement for FAO, a simpler symbol is FAO for the respiratory activity with FAO-linked substrates. (ii) 'CI+ETF' indicated respiration in the presence of a cocktail of Complex I-linked substrates (glutamate&malate) and electron transferring flavoprotein-linked substrate (octanoylcarnitine). It is suggested to use the symbol FN (or FAO&CI) to indicate respiration measured in the presence of such a F-junction and N-junction substrate cocktail, which then is distinguished from FAO+CI which is the calculated sum of respiratory activities measured separately with F- and N- (FAO-linked and CI-linked) substrates. Similarly, 'CI+II+ETF' is replaced by FNS (FAO&CI&II).
O2k-Publications
Bioblast Alert
- The most popular O2k-Publication on Bioblast of the year 2013 and 2014.
- Bioblast alert 2011 (4)